#SpatialTranscriptomics
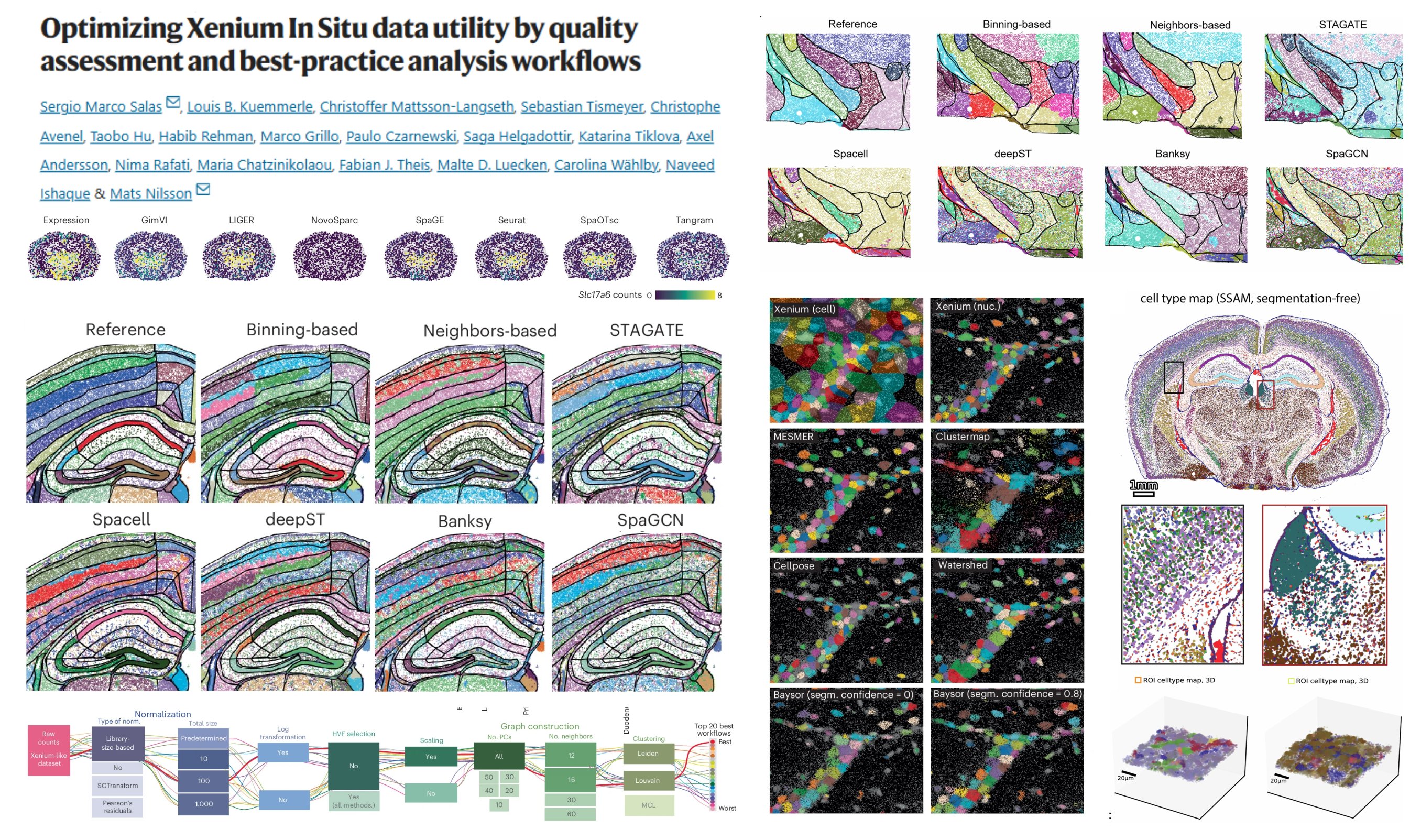
Optimizing Xenium
@10xGenomics In Situ data utility by quality assessment & best-practice analysis workflows
If you are a Xenium user, this is a must-read🔥
👉Segmentation: Cellpose + Baysor (Segmentation-free alternative: SSAM, Points2Regions)
👉Preprocessing: Library size normalization, log-transformation, scaling, k-nearest neighbors graph (all PCs + 16 neighbors), Louvain clustering
👉Spatially variable feature selection: Squidpy Moran's I, Seurat Moran's I, Sinfonia, SomDE, Giotto, Hotspot
👉Imputation: SpaGE, Seurat, Tangram, SpaOTsc
👉Spatial domain identification: binning-based strategy, Bansky, SPACEL
Performance
vs CosMX, Molecular cartography, MERFISH, MERSCOPE, high-sensitivity ISS
@naturemethods 2025
nature.com/articles/s4159…